Antonio Jimenez-Marin, Nele De Bruyn, Jolien Gooijers, Alberto Llera, Sarah Meyer, Kaat Alaerts, Geert Verheyden, Stephan P. Swinnen, and Jesus M. Cortes. Multimodal lesion network mapping for prediction of sensorimotor behavior in stroke patients. OHBM 2022 – Organization for Human Brain Mapping [pdf]
Introduction:
Mapping the behavioral impact of brain lesions is of crucial importance for clinical practice. Regardless of the cause of the lesion, its size or the precise location, accumulated evidence in recent years has pushed forward the connectome hypothesis to show that the belonging of a lesion to a specific network can predict a multitude of responses in the patient across behavioral, cognitive, or sensory-motor domains. Lesion network mapping (LNM) is a successful technique for mapping symptoms to neural networks after acquired brain injury. Beyond lesion characteristics such as its etiology, size or location, LNM has shown that when a subgroup of patients has common symptoms after an injury this is because their lesions are part of the same network, thus linking symptoms to specific networks. Here, we extend LNM according to two strategic avenues. Firstly, by proposing a multimodal approach in which we introduce a combination of both structural and functional networks to predict behavior. Secondly, by assessing behavioral performance using a combination of several multi-domain scores. For this purpose, we employed a canonical correlation analysis to link multi-domain behavior to different lesion connectivity maps.
Methods:
We analyzed N=54 first-stroke patients (25 males) with a mean age of 68.78 (σ = 13.98; range 28 – 92 years). Every patient was assessed with the same four tests: Action Research Arm Test (ARAT) and Fugl-Meyer assessment – upper extremity (FMA-UE) as motor variables; and Erasmus-modified Nottingham Sensory Assessment (Em-NSA) and perceptual threshold of touch (PTT) as somatosensory variables.
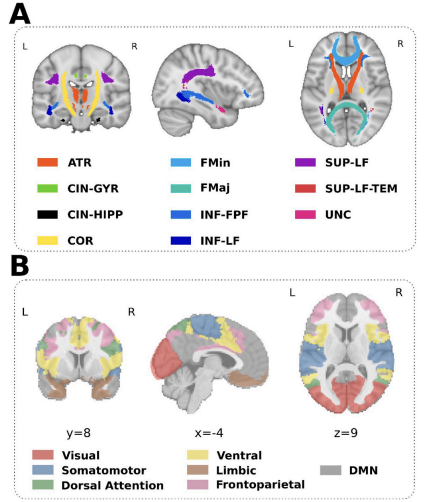
We build structural connectivity (SC) and functional connectivity (FC) maps of each patient-lesion mask using data of the Human Connectome Project with 1000 healthy participants. We reduced the dimension of the lesion connectivity maps using Principal Component Analysis (PCA) and associated the multimodal imaging components to the sensory-motor scores applying a Canonical Correlation Analysis (CCA). In order to overcome the overfitting, we performed leave-one-out cross-validation. The methodology was performed for SC and FC separately (unimodal analyses) and combined by spatial concatenation of the SC and FC matrices (multimodal analysis), followed by PCA and using the obtained mixed components to apply CCA similarly as we did for the unimodal cases.
Results:
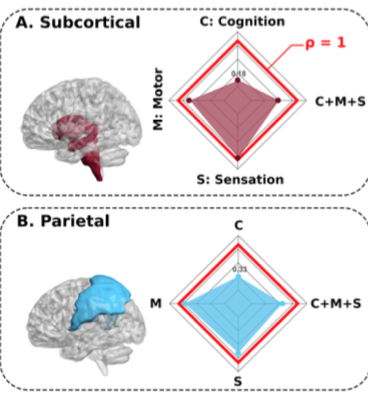
Our results are three-fold. Firstly, the multimodal analysis reveals that functional connectivity maps contributed more than structural connectivity maps in the optimal prediction of sensorimotor behavior. Secondly, the maximal association solution between behavioral outcome and multimodal lesion connectivity maps provided equally contribution of sensory and motor coefficients, in contrast to the unimodal analyses where the sensory contribution dominated in both structural and functional maps. Finally, when looking at each modality individually, the performance of the structural connectivity maps strongly depended on whether sensorimotor performance was corrected for lesion size (thus eliminating the effect that larger lesions produce higher sensorimotor dysfunction). In contrast, the maps of functional connectivity provided similar performance with and without correcting for lesion size.
Conclusions:
The application of our multimodal LNM approach to predict multidomain sensorimotor behavior, supports the synergistic and additive features that different types of brain networks have on patient’s outcome after brain injury, thereby making the whole more than the sum of its parts. Moreover, when the patient’s behavior is assessed across multiple domains of cognitive, sensory and motor function, our methodology combining structural and functional maps appears most suitable when assessing such multidomain outcomes.
References
[1] Aerts H, Fias W, Caeyenberghs K, Marinazzo D (2016): Brain networks under attack: robustness properties and the impact of lesions. Brain 139:3063–3083.
[2] Baldassarre A, Ramsey LE, Siegel JS, Shulman GL, Corbetta M (2016): Brain connectivity and neurological disorders after stroke. Curr Opin Neurol 29:706–713.
[3] Boes AD, Prasad S, Liu H, Liu Q, Pascual-Leone A, Caviness VS, Fox MD (2015): Network localization of neurological symptoms from focal brain lesions. Brain 138:3061–3075.
[4] Corbetta M, Ramsey L, Callejas A, Baldassarre A, Hacker CD, Siegel JS, Astafiev SV, Rengachary J, Zinn K, Lang CE, Connor LT, Fucetola R, Strube M, Carter AR, Shulman GL (2015): Common Behavioral Clusters and Subcortical Anatomy in Stroke. Neuron 85:927–941.
[5] Darby RR, Joutsa J, Fox MD (2019): Network localization of heterogeneous neuroimaging findings. Brain 142:70–79.
[6] Fox MD (2018): Mapping Symptoms to Brain Networks with the Human Connectome. N Engl J Med 379:2237–2245.
[7] Griffis JC, Metcalf NV, Corbetta M, Shulman GL (2020): Damage to the shortest structural paths between brain regions is associated with disruptions of resting-state functional connectivity after stroke. NeuroImage 210:116589.
[8] Ramsey LE, Siegel JS, Lang CE, Strube M, Shulman GL, Corbetta M (2017): Behavioural clusters and predictors of performance during recovery from stroke. Nat Hum Behav 1:0038.
[9] Salvalaggio A, De Filippo De Grazia M, Zorzi M, Thiebaut de Schotten M, Corbetta M (2020): Post-stroke deficit prediction from lesion and indirect structural and functional disconnection. Brain 143:2173–2188.
[10] Siegel JS, Ramsey LE, Snyder AZ, Metcalf NV, Chacko RV, Weinberger K, Baldassarre A, Hacker CD, Shulman GL, Corbetta M (2016): Disruptions of network connectivity predict impairment in multiple behavioral domains after stroke. Proc Natl Acad Sci 113:E4367–E4376.