Peter beim Graben, Antonio Jimenez-Marin, Ibai Diez, Jesus M. Cortes, Mathieu Desroches and Serafim Rodrigues. Metastable Resting State Brain Dynamics. Frontiers in Computational Neuroscience 13:62, 2019 [pdf]
Abstract
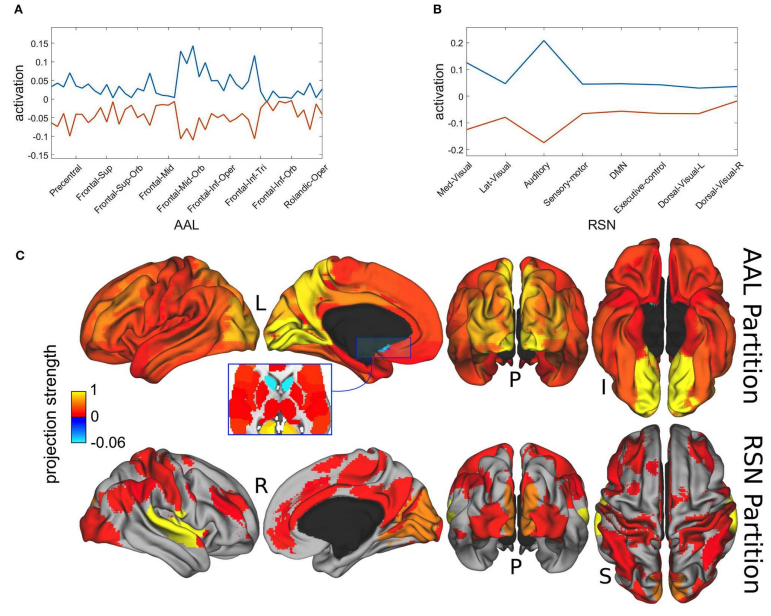
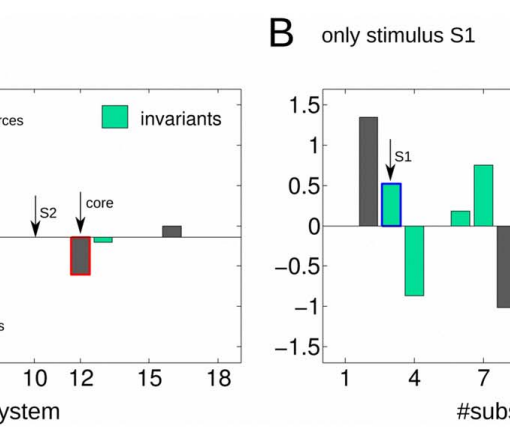
Metastability refers to the fact that the state of a dynamical system spends a large amount of time in a restricted region of its available phase space before a transition takes place, bringing the system into another state from where it might recur into the previous one. beim Graben and Hutt (2013) suggested to use the recurrence plot (RP) technique introduced by Eckmann et al. (1987) for the segmentation of system’s trajectories into metastable states using recurrence grammars. Here, we apply this recurrence structure analysis (RSA) for the first time to resting-state brain dynamics obtained from functional magnetic resonance imaging (fMRI). Brain regions are defined according to the brain hierarchical atlas (BHA) developed by Diez et al. (2015), and as a consequence, regions present high-connectivity in both structure (obtained from diffusion tensor imaging) and function (from the blood-level dependent-oxygenation—BOLD—signal). Remarkably, regions observed by Diez et al. were completely time-invariant. Here, in order to compare this static picture with the metastable systems dynamics obtained from the RSA segmentation, we determine the number of metastable states as a measure of complexity for all subjects and for region numbers varying from 3 to 100. We find RSA convergence toward an optimal segmentation of 40 metastable states for normalized BOLD signals, averaged over BHA modules. Next, we build a bistable dynamics at population level by pooling 30 subjects after Hausdorff clustering. In link with this finding, we reflect on the different modeling frameworks that can allow for such scenarios: heteroclinic dynamics, dynamics with riddled basins of attraction, multiple-timescale dynamics. Finally, we characterize the metastable states both functionally and structurally, using templates for resting state networks (RSNs) and the automated anatomical labeling (AAL) atlas, respectively.