Sebastaino Stramaglia, Javier Rasero, Jesus M. Cortes, Daniele Marinazzo, Ibai Diez. Connectome sorting by Consensus Clustering increases separability in group neuroimaging studies. OHBM 2019 – Organization for Human Brain Mapping [pdf]
Introduction:
A fundamental challenge in preprocessing pipelines for neuroimaging datasets is to increase the signal-to-noise ratio for subsequent analyses. This can be achieved for example by getting more compact groups before performing any statistical comparison. In this regard, the consensus clustering algorithm recently developed for brain connectivity data (Rasero et al. 2017) copes with the heterogeneity of classes of subjects as defined by phenotypic labels (e.g. healthy and patients). As a consequence, we suggest (Rasero et al. 2018) that the application of the consensus clustering approach to neuroimaging studies can be a valid additional step for connectome processing to find subgroups of subjects with reduced intra-group variability and therefore increasing the separability of the distinct subgroups when connectomes are used as a biomarker. We also demonstrate that unique regions within each cluster arise, information which is not identified when the groups are used as a whole, bringing new insight that could be relevant from a clinical point of view.
Methods:
The consensus clustering is summarised as follows: (i) for each brain region’s pattern connectivity, a distance matrix is computed (ii) cluster each distance matrix using k-medoids clustering algorithm, (iii) build the modularity matrix as the consensus network from the corresponding partitions subtracting the null ensemble of co-assignments. Consequently, this method is applied individually to Typically Developing and Autsim Spectrum Disorder (ASD) groups in a cohort of 150 subjects from the ABIDE initiative in order to extract the natural
classes presented in the data. A subsequent supervised analysis by means of a multivariate distance matrix regression is performed between the subgroups found, computing the increase of separability between both classes as measured by the coefficient of determination “pseudo r-squared” and identifying unique regions whose pattern is altered only in the subgroups.
Results:
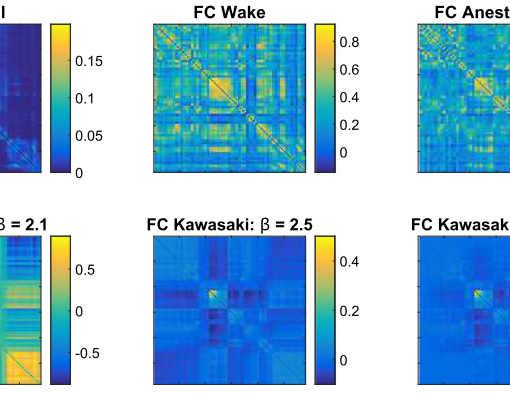
Individual application of the consensus clustering algorithm to both groups yields a modularity matrix with 3 communities in the TD group and a modularity matrix with 2 clusters in the ASD group. Moreover, a Wilcoxon Rank sum test performed on the ASD community partition separates subjects statistically by age (p=0.0028) and framewise displacement (p<0.0001). Likewise, after group partition most of the brain regions exhibit altered pattern connectivities which amplify the class labelling effect size. This is translated into a gain of separability that allows to unmask regions not observed significantly different when whole groups are taken together. For example, when ASD partitions are used in group comparison with TD subjects, the first cluster elucidates unique regions located in the Inferior Frontal Gyrus, the Frontal Pole, the Precuneus, Postcentral and Paracentral areas. On the other hand, the second cluster displays unique regions belonging to areas in the Superior Temporal Sulcus, the Superior Temporal Lobe, the Inferior Temporal Lobe and the Supramarginal gyrus.